Rho-associated coiled-coil containing kinase 2 (ROCK2), a large, coiled-coil containing, membrane anchored, sophisticated protein plays important role in many diseases including cardiovascular, Alzheimer's, Parkinson's diseases, and cancer. ROCK2 is assumed to play an important role in the plasticity of cancer cell migration, the phenomenon which secures survival advantage to the cancer cells during drug treatments. Pharmaceutical companies make significant effort to develop ROCK2 inhibitors for treating various diseases, including drugs that have potential to prevent cancer from spreading by blocking cell migration. Inhibitor development is difficult because of the lack of ROCK2 structure and without knowing the structure-based mechanism of function.
ROCK kinases are localized mainly in the cytoplasm with their C-terminal domains anchored to the cell membrane. As the locations of the downstream targets differ by as much as 100 nm, the conformation of the active ROCK proteins must sample a large conformational space. In their native form, ROCK kinases are enzymatically inactive due to auto-inhibition of the N-terminal kinase domain by the distant C-terminal region.
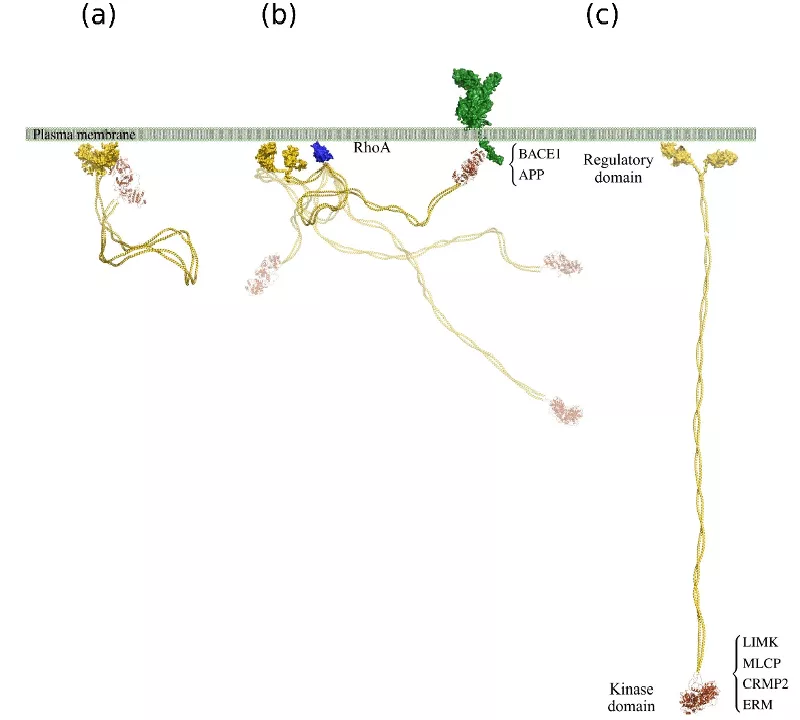
ROCK2 is anchored to the membrane by its C-terminal domain, and the N-terminal kinase domain is folded back forming a compact conformation with limited flexibility (a). Since the binding between the terminal domains is weak, a substrate capable of forming a strong interaction with the kinase domain can displace the C-terminal domains. Binding of RhoA induces transient loosening of the compact structure, and releases the N-terminal part of the molecule (b), allowing interactions with substrates at distant locations, e.g. the cytoskeleton. Such interactions stabilize the extended conformation of ROCK2 (c).
Since the activity of ROCK2 is essential for several normal physiological functions, direct inhibition of the kinase activity cannot be applied in drug development because of the expected severe side effects.
Researchers at the ELKH Research Centre for Natural Sciences, succeeded in expressing this large, complex, multidomen, dimeric protein in quantities sufficient for structural studies. This has opened the way for a group of scientists, led by Péter Závodszky to reveal the solution structure of this membrane anchored long flexible protein, and the localization and mapping of potential allosteric functional sites that can be targeted for switching of protein-protein interactions responsible for disease, while leaving the essential kinase activity intact.
The present solution small-angle X-ray scattering study revealed that ROCK2 population is a dynamic mixture of folded and partially extended conformers (Figure). It has also been shown that the binding of the natural protein substrate, RhoA to the coiled-coil domain shifts the equilibrium towards the partially extended state. Enzyme activity measurements suggest that the binding of other (LIM kinase 1 and 2, myosin phosphatase) natural protein substrates to the kinase domain breaks up the interaction between the N-terminal kinase and C-terminal regulatory domains, but binding of smaller substrate analogues does not. The present study reveals the dynamic behavior of this long, dimeric molecule in solution, and our structural model provides a mechanistic explanation for a set of membrane-proximal functions while allowing for the existence of an extended conformation in the case of membrane-distal functions. This structural information opens the way for designing drugs, selective for the erroneous functions while leaving essential functions intact. The paper was recently published in Nature Communications Biology (“Ligand-induced conformational rearrangements regulate the switch between membrane-proximal and distal functions” by István Hajdú, András Szilágyi, Barbara M. Végh, András Wacha, Dániel Györffy, Éva Gráczer, Márk Somogyi, Péter Gál and Péter Závodszky)